Next-Gen Protein Sequencing Made Scalable, Accessible, and Insightful
The next frontier in discovery is set to transform our understanding of the human proteome and unlock the intricacies of cellular function like never before.
Next-gen protein sequencing (NGPS) reveals insights into critical questions:
- Is a specific protein present?
- Which proteins are important to health and disease?
- Why are proteins important?
- How do proteins change with disease and in response to therapy?
With NGPS, you can answer these critical questions quickly, right in your own lab.

Learn About Next-Gen Protein Sequencing Technology
Benchtop-friendly and accessible for all labs
Auto-analysis eliminates the need for bioinformatics
Single-molecule resolution for deeper insights
Streamlined setup with minimal maintenance
How it Works
Prepare
Library Preparation and Sequencing Kits contain everything you need to initiate the next-gen protein sequencing process, from sample prep to loading.
- Includes reagents to digest and functionalize proteins
- Supplies aminopeptidases and recognizers to begin sequencing
Sequence
Platinum® Pro sequences peptides by capturing fluorescent signals from each N-terminal amino acid (NAA) binding event, enabling real-time, single-molecule protein sequencing.
- Uses aminopeptidases to sequentially cleave each amino acid
- Repeats the process to sequence the entire peptide
Analyze
Sequencing data is automatically uploaded to the Platinum Analysis software, which interprets results at the single-molecule level. No bioinformatics expertise is required.
- Delivers clear, interpretable single-molecule protein insights
- Removes the need for complex data analysis tools
Identify Peptides with Kinetic Signatures
Accelerating the Path to Discovery
A kinetic signature comprises the measurable characteristics of the series of dynamic recognizer events that uniquely identify a peptide.
Kinetic signatures change dynamically with alterations in amino acid and post-translational modifications (PTMs), providing a reliable means to accurately align peptides to their respective proteins.
Unlike other technologies, kinetic signatures offer a robust and confident approach to this alignment.
As next-gen protein sequencing becomes more widespread, kinetic signatures will be crucial to identifying every amino acid and every PTM, making Quantum-Si’s technology key to advancing our understanding of the proteome.
Contact Our Experts
How NGPS Compares to Other Methods
NGPS Takes Mass Spectrometry to New Depths
Our technology identifies proteins using unique kinetic signatures based on the binding kinetics of amino acid recognition events. In contrast, mass spectrometry infers protein identification based on the mass:charge ratio, which may not distinguish protein variants of similar size.
| Mass Spectrometry | Platinum | |
|---|---|---|
| Expertise | Advanced | Basic |
| Analysis | Complex | Automated |
| Lab Space | Designated Space | Benchtop Instrument |
| Startup Cost | $$$$$ | $ |
| Maintenance Cost | $$$ | $ |
Next-gen protein sequencing with Platinum Pro fits your lab’s space, budget, and expertise, delivering rapid insights without the need for bioinformatics or complex infrastructure.
NGPS Improves Upon Immunoassays
Next-gen protein sequencing provides deeper insights with single-molecule resolution that, unlike immunoassays, enables assessment of what is happening at the amino acid level.
- Verify the specificity of your antibodies by sequencing the target protein
- Sequence the protein band from your gel
- Positively identify protein findings with amino
acid resolution - Interrogate protein variants not resolved by antibodies
NGPS Improves Upon Edman Degradation
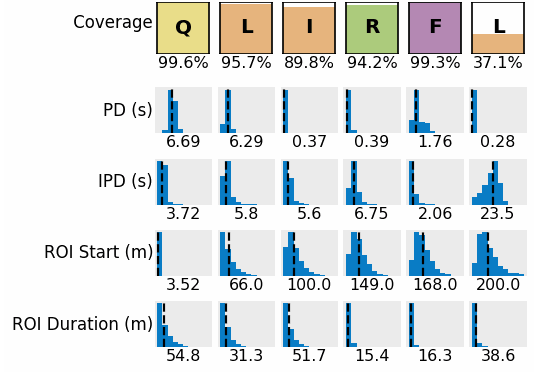
Next-gen protein sequencing leverages a one-pot reaction without complex and expensive cyclical chemistry and fluidics to achieve single-amino acid resolution. With NGPS, you benefit from:
- Deeper protein sequencing discoveries
- Easy protein sequencing workflows
- Affordable protein sequencing solutions
Explore the Capabilities of Platinum Pro
Discover how our proteomics platform can transform your research with single-molecule resolution and unmatched flexibility.
“This paper highlights the potential of merging next-generation protein sequencing with individual ion mass spectrometry. By combining these methods, we can capture a fuller picture of IL-6 proteoforms, which is critical for advancing therapeutic innovation.”
Neil Kelleher, PhD
Professor, Northwestern University
Next-Gen Protein
Sequencing Applications
Protein Barcoding
Simultaneously characterize multiple protein variants with barcoding, increasing throughput and improving efficiency in screening and therapeutic development.
Protein Identification
Achieve single-amino acid resolution for precise protein characterization, and validation in research and diagnostics.
Protein Variants
Detect amino acid substitutions and isobaric variations with single-molecule resolution, uncovering critical insights into health and disease.
Post Translational Modifications
Precisely analyze proteoforms and post-translational modifications (PTM) to gain deeper functional insights into protein regulation and activity.
Antibody Sequencing and Characterization
Gain deep insights into antibody specificity, affinity, and purity. This advanced approach ensures precise targeting and maximized efficacy in therapeutic development.